Welcome to the Nanocluster Interpolation Scheme Program (NISP) documentation!¶






Section author: Geoffrey Weal <geoffrey.weal@gmail.com>
Section author: Dr. Anna Garden <anna.garden@otago.ac.nz>
Section author: Dr. Andreas Pedersen
Section author: Prof. Hannes Jónsson <hj@hi.is>
Group page: https://blogs.otago.ac.nz/annagarden/
Page to cite with work from: A. L. Garden, A. Pedersen, H. Jónsson, “Reassignment of ‘magic numbers’ of decahedral and FCC structural motifs”, Nanoscale, 10, 5124-5132 (2018), DOI: 10.1039/C7NR09440J
What is this Documentation about?¶
This documentation is designed to guide the user to use the Nanocluster Interpolation Scheme Program (NISP) program.
What is NISP¶
The NISP program is an interpolation scheme that is designed to give an approximate guide for the estimated energies of unsymmetric nanoclusters based on energetic trends between perfect, closed-shell nanoclusters. This program will create all the perfect, close shell icosahedral, decahedral, and octahedral clusters that can be created between 13 atoms and an upper atom number limit. These nanoclusters are locally optimised using either an ASE, an ASE-integrated calculator, or with VASP. After the nanoclusters are locally optimised, the delta energy is obtained for each nanocluster before providing plots and text files that indicate the estimated energies of perfect, closed-shell and unsymmetric nanoclusters and how to remove atoms from the larger perfect closed-shell nanocluster to give unsymmetric nanoclusters with a certain number of atoms. See “Reassignment of ‘magic numbers’ of decahedral and FCC structural motifs” (DOI: 10.1039/C7NR09440J) for more information about how this interpolation scheme works.
The algorithm was designed by Dr Anna Garden of the University of Otago, Dunedin, New Zealand, and Dr. Andreas Pedersen and Prof. Hannes Jónsson of the University of Iceland. The Github page for this program can be found at github.com/GardenGroupUO/NISP.
Dr. Anna Garden: blogs.otago.ac.nz/annagarden
Dr. Andreas Pedersen: https://dk.linkedin.com/in/andreas-pedersen-a847025
Prof. Hannes Jónsson: english.hi.is/staff/hj
Try Organisms before you Clone/Pip/Conda (on Binder/Jupter Notebooks)!¶
If you are new to the NISP program, it is recommended try it out by running NISP live on our interactive Jupyter+Binder page before you download it. On Jupyter+Binder, you can play around with the NISP program on the web. You do not need to install anything to try NISP out on Jupyter+Binder.
Click the Binder button below to try NISP out on the web! (The Binder page may load quickly or may take 1 or 2 minutes to load)
Installation¶
It is recommended to read the installation page before using the NISP program. See Installation: Setting Up NISP and Pre-Requisites Packages for more information. Note that you can install NISP through pip3 and conda.
Output files that are created by NISP¶
An example of the plots that are created is the interpolation scheme plot, which shows all the estimated energies of nanoclusters across the size range of nanoclusters that you are measuring across. An example of this for Au nanoclusters, using the RGL potetial with parameters from Baletto et al. (DOI: 10.1063/1.1448484), is shown below:
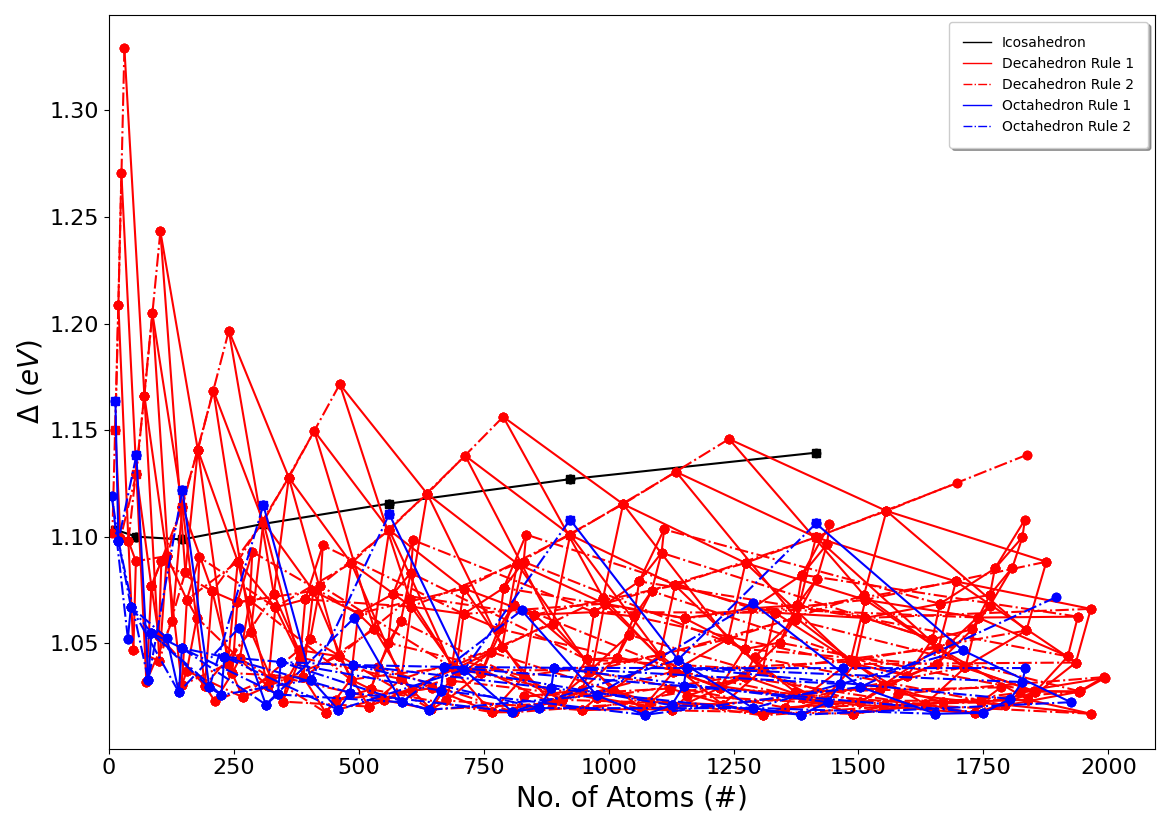
There are also other plots created by NIPS as well as other text documents that contain the delta energies of the various nanoclusters that you calculated, as well as instructions about how to remove atoms from certain nanoclusters in order to get icosahedral, decahedral, and octahedral nanoclusters with the particular number of atoms that you desire. Click here to see examples of all of these plots and text files.
Table of Contents¶
- How the Nanocluster Interpolation Scheme Program (NISP) works
- Installation: Setting Up NISP and Pre-Requisites Packages
- Interpolation_Script.py - How to run NISP
- RunMinimisation.py - Writing a Local Minimisation Function for NISP
- How to perform NISP with VASP calculations
- How to manually enter energy results into NISP
- How to obtain cohesive energies
- What are delta energies?
- Examples of Running NISP with Interpolation_Script.py
- Examples of data that the NISP program gives
- Helpful Programs to run NISP
- NISP Python Files
- Index
- Python Module Index